Session Overview¶

This view shows all the scans in a session, automatically imports data from the host AFM, sorts out unwanted scans to a sub-folder, adjust images for synchronization problems with the host AFM, and generates parameter maps on multiple scans in a batch processing mode.
Session¶
Open IMP session and choose a session folder. An overview is generated showing file info and image thumbnails where each row corresponds to one scan. Reload causes the overview to be updated to the current state of the session folder. View session log opens a window showing the session log file.
Import host AFM data¶
The AFM height image is a record of the change of the probe height during the x-y scan. The probe height 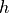 , is the position of the base or the fixed end, not the position of the tip
, is the position of the base or the fixed end, not the position of the tip  at the free end of the flexing cantilever (see drawing in the section Force Reconstruction Methods). The probe height is controlled by the scanning feedback and in most modern AFMs the height signal is measured using a linear sensor built in to the scanner. When performing ImAFM™, the height data is recorded by the host AFM. To associate ImAFM™ quantitative analysis with this height data, it is necessary to import the height data in to the IMP Software suite.
at the free end of the flexing cantilever (see drawing in the section Force Reconstruction Methods). The probe height is controlled by the scanning feedback and in most modern AFMs the height signal is measured using a linear sensor built in to the scanner. When performing ImAFM™, the height data is recorded by the host AFM. To associate ImAFM™ quantitative analysis with this height data, it is necessary to import the height data in to the IMP Software suite.
Associate host AFM files
Import height data: Navigate to the folder on your host AFM computer, network or storage device where the host AFM files are stored. When you OK the selected folder theMatch Imageswindow opens. Drag the image thumbnails fromUnmatched IMP imagesat the bottom, and drop them on the correspondingAFM imagesat the top. When you have matched a few images, clickMatch with timestampsand the computer will attempt to associate all remaining files. If the matching is incorrect, you can drag images away from middle group ofMatched IMP images. When the matching looks correct, clickOKand the AFM height data will be appended to the .imp file. (Note: In DI/Veeco Nanoscope software version 5 there is a bug which requires you to simply open and close the file in the DI software before you can import it to the IMP suite.)Hide uncheckedwill move all the unchecked scans to a sub-folder of the session folder called ‘hidden_scans’. These scans will no longer be displayed in the session overview, but they are not erased. You can always use your computers file browser to move a file from the ‘hidden_scans’ folder back to the session folder, where it will again be displayed when you clickReload.Unhide allbrings all files from the ‘hidden_scans’ sub-folder up to the session folder.
Flip and swap scans¶
Because the host AFM does not code the trigger signals for trace and retrace (only for change of fast-scan direction) it may be necessary to left/right flip the scan data or exchange the trace and re-trace data in order to associate the ImAFM™ scan correctly with the host AFM scan. Furthermore, AFM triggers do not code for slow scan direction (only end of frame) so it may be required to exchange up/down flip images. All this is easily done in the Session Overview. To perform these these flip and swap operations you first have to select the scans.
Checkmarks control which files will be flipped and swapped: Check will select all scans, Uncheck deselects all scans, Toggle will switch checked to unchecked, and unchecked to checked.
Apply to checked scans
Flip Left/Rightis performed on both trace and retrace data, but trace and re-trace are not exchanged.Flip Up/Downis performed on both trace and retrace data, but trace and re-trace are not exchanged.Swap Trace/Retracewill exchange trace and retrace, with out performing any flip. Note that left-right flip, does not necessarily imply that trace and retrace were stored incorrectly when scanning. Depending on whether you selected the trace or retrace for storing the height data in the host AFM, you may want to exchange trace an retrace to get correct association. ImAFM™ always stores both trace and retrace.
Batch process parameter maps and force volume data¶
Analyze checked scans selects which type of Parameter Maps will be made on the checked scans in a batch process. As parameter maps are computationally demanding, it is very useful to set up and run batch processes.
Parameter Map: Model fit, when checked, will perform the analysis described in Model Fit, generating parameter maps, or color coded images of tip-surface force parameters. Thesettingsbutton opens a panel for choosing the model and fit parameters, as described in Parameter Maps.Parameter Map: Polynomial, when checked, will perform the analysis described in Fast Polynomial, based on the polynomial representation of the conservative force-distance curve. Thesettingsbutton opens a dialog with options similar to that for Parameter Maps.Polynomial degreeis described in Polynomial.ADFS Force Volume, when checked, will perform Amplitude Dependent Force Spectroscopy (ADFS) on each pixel of an image. Thesettingsbutton opens a dialog with options similar to that for Parameter Maps.Save to HDF5will create one file containing an ADFS force curve at each pixel, whereasSave to ASCIIwill create a folder with one file for each pixel, where the pixel coordinates are given in the file name.Startthe batch processing for all checked scans. A status bar tells you how far the batch process has progressed.Abortwill end the process, storing the data analyzed thus far. It is not possible to resume an aborted process.
Three dimensional viewer¶
To generate a three dimensional view you must have imported the height data from your host AFM. Double click on the height image to open the IMP 3D-viewer. Left click and drag on the image to rotate the viewing angle. Right click and drag to change the zoom.
The Viewer settings group allows for change of the 3D image properties, including the Z scale which usually defaults to a factor larger than 1, in comparison to the X and Y scales. Such scaling exaggerates the topography.
The Texture settings group controls which quantity is represented by the color painted on the topography. Default is the response phase at the first drive frequency. You can choose to plot Amplitude or Phase of any frequency in the intermodulation spectrum. If the .imp file has an Add-on image, for example an ImEFM™ image or a parameter map, these can also be painted on the topography. You can also choose to a Parameter file consisting of 2D arrays, the same size as the image, stored as a numpy .npz file. Limits opens up a dialog box with a histogram of image pixels, allowing you to set the upper and lower limits of the color scale.
The Camera position is shown in the boxes. It is easiest to adjust this by left or right click-and-drag on the image. If the Animate box is checked, the image will rotate around the azimuth. You can use screen capture programs to make a movie of this rotation.
The Plane fit group has three different methods for flattening the image:
Noneshows the height data as it is stored in the host AFM height image file.
Averagefits a plane over the entire image and offsets all height data so that this plane is flat.
Selected pointsallows you to flatten on a selected region of the image. With this option selected, hover over the image with the mouse. When you press the ‘p’ key, the point at the mouse arrow will be selected. After selecting 3 points a plane can be calculated and the image will offset all height data so that this plane is flat. You can continue to select more than 3 points, and the image will be flattened with a best-fit plane to all selected points. Theclearbutton removes all selected points. A red frame appears when you select points. To make this frame disappear, hover outside the image and press the ‘p’ key.
Save image as... opens a save dialog so you can save to a desired location, and choose between several different file formats.
Render movie will create a movie of a single rotation around the azimuthal angle, as see in the viewer when the Animate box is checked. The movie uses 3D ray tracing so it takes a rather long time to generate the movie. This feature is experimental and it requires that you install Povray software.
Render 3D will use a 3D ray tracing program to create the same view seen in the viewer, with shadows added. It can take several seconds to render the image. The image will be stored in your session folder, in a sub-folder called Povray. This feature is experimental and it requires that you install Povray software.
